2020-03-15 11:31:13
Chao Lu 1, Ting Wen 1,*, Maochao Zheng 3, Daojun Liu 3, Guilan Quan 1,2, Xin Pan [ Introduction
Introduction
In the past decades, nanotechnology has been well developed to construct drug delivery systems, including but not limited to micelles, liposomes, and nanoparticles [1-10]. Compared with conventional formulations, these nanoscaled drug delivery systems (DDS) exhibit great potential in improving drug stability, preventing premature drug release, altering drug distribution, and prolonging the half-life of drugs [11]. Therefore, they are widely applied in the delivery of a great variety of drugs including anticancer drugs [1,2], antimicrobial agents [3,4], and anti-inflammatory drugs [5,6], among others.
Though showing unique advantages in delivering drugs for monotherapy, the nanoscaled DDS still suffers from many limitations in enhancing the comprehensive treatment efficacy of disease with multiple etiologies. In fact, many common clinical diseases are caused or worsened by the interplay of multiple factors. For example, burn wound infection is often aggravated by the infection of bacteria, such as Pseudomonas aeruginosa (P aeruginosa) and Staphylococcus aureus (S. aureus) [12]. When a massive loss of liver parenchyma and metabolic function induces acute liver failure, patients will become susceptible to pathogens, which largely limits the feasibility of advanced emergency liver transplantation [13]. It was also reported that bacterial infection might be one of the main reasons to induce tumour formation, so both anticancer and antibacterial measures should be employed for the treatment of these diseases, such as Helicobacter pylori-induced gastric carcinoma [14]. Therefore, to treat multifactorial diseases, it is critical that DDS should be rationally developed to co-deliver therapeutic agents and antimicrobial agents such as antibiotics [3], antimicrobial peptides [15,16], metal ions [4], and natural compounds [17].
To achieve this, one of the most common strategies is to co-deliver two or more drugs in one formulation [18,19]. However, due to limited drug loading space of nanocarriers, the co-delivery strategy usually fails to achieve sufficient loading of multiple drugs simultaneously [18-20]. Recently, poly(i-lysine) (PLL)-based polymers have been fabricated to encapsulate drugs with a negative charge, such as small molecule drugs [21-23], proteins [24,25], and nucleic acids [26,27]. Notably, Qiao et al. [15] found that PLL-based polymers exerted a bactericidal mechanism similar to that of common antimicrobial peptides (AMPs), possessing a great potential to eradicate bacterial infections. Our previous research [16] also demonstrated that multi-armed PLL (MPLL), with high antimicrobial activity, enormous selectivity, and remarkable proteolytic stability, represented a new series of potent antimicrobial peptides to treat drug-resistant infections [16]. All these research advancements prognosticated that PLL-based polymers could be served not only as drug carriers but also as macromolecules with therapeutic activity to synergize therapeutic effect, which implied a new perspective for the design of nanomedicine against multifactorial diseases.
In this research, poly (ethylene glycol) (PEG) crosslinked alternating copolymers in the form of MPLL-alt-PEG (Scheme 1) were designed, synthesized, and evaluated as a platform for both anionic drug delivery and anti-bacteria. MPLL with a branched polyethyleneimine (PEI) core and poly(i-lysine) peripheral chains comprised the polyelectrolyte segment of MPLL-alt-PEG copolymer. Their multi-armed molecular architecture possessed high cationic charging density, which is beneficial for enhancing the binding affinity towards anionic drugs and bacterial lipid bilayers via electrostatic interactions [16]. It is expected that MPLL-alt-PEG could encapsulate anionic drugs, followed by self-assembly into shell crosslinked polyion complex (PIC) micelles (Scheme 1). Our previous research suggested that shell crosslinked micelles could possess unique advantages ineffective drug loading, improving micellar stability, and protecting the drug from premature release and degradation [28]. Meanwhile, the antimicrobial activity of MPLL could also be improved after crosslinking since the crosslinking reaction would lead to increased molecular weight, proteolytic stability, and in vivo half-life [16,29]. The drug-loaded MPLL-alt-PEG micelles could integrate the therapeutic effect of loaded drugs and the antimicrobial activity of carriers. We demonstrated the feasibility of MPLL-alt-PEG to serve as a DDS for small molecule drugs and therapeutic biomacromolecules, using water-soluble methyl orange (MO) and immunotropic agent recombinant human interferon-a-2b (IFN) as anionic model drugs. Subsequently, the antimicrobial activity and mechanism of MPLL-alt-PEG was investigated using Gram-positive methicillin-resistant Staphylococcus aureus (MRSA) and Gram-negative P. aeruginosa as model bacteria. We expected that the MPLL-alt-PEG-based nanoplatform could be applied as a potential system to treat bacterial infection-involved multifactorial diseases.
2.Materials and Methods
2.1.Materials
Branched PEI (Mw = 0.8 kDa) was obtained from Sigma-Aldrich (Shanghai, China). g-Benzyloxycarbonyl-L-lysine (ZLL) was purchased from GL Biochem (Shanghai, China) and used without further purification. PEG (succinimidyl carboxymethyl ester)2 (SCM-PEG-SCM, Mw = 7.5 kDa) was supplied by byJenKem Technology Co., Ltd. (Beijing, China). Methyl orange, anisole, methanesulfonic acid, trifluoroacetic acid (TFA), and fluorescein isothiocyanate (FITC) were all obtained from Aladdin (Shanghai, China). A dialysis tube with molecular weight cut-off (MWCO) values of 7 and 14 kDa was produced by Viskase (Darien, IL, USA), while dialysis tubes of 50 kDa and 100 kDa were purchased from Spectrum Laboratories Inc. (Los Angeles, CA, USA). IFN (5 million U) was obtained from Anhui Anke Biotechnology Co., Ltd. (Anhui, China).
2.2.Synthesis and Characterization of MPLL-alt-PEG Copolymers
MPLL-alt-PEG copolymers were synthesized in a successive procedure of PEI initiated polymerization of e-benzyloxycarbonyl-L-lysine N-carboxy anhydrides (ZLL-NCA) to prepare PEI-grafted poly(e-benzyloxycarbonyl-L-lysine) (PEZ), followed by the peripheral crosslinking with difunctional SCM-PEG-SCM, and the subsequent removal of the benzyloxycarbonyl side-chain protecting groups (Figure 1).
In the first step, ZLL-NCA was prepared via phosgenation of ZLL in anhydrous ethyl acetate (Yield: 78%) and then polymerize d using PEI as the macroinitiator to obtain the star-block copolymers PEZ (Yield: 92%) according to our previous report [16,24,28,30].
Then, 100 mg of PEZ was dissolved in a mixture of 50 mL of DMSO and 250 mL of dichloromethane (DCM). The solution of SCM-PEG-SCM at a concentration of 0.2 g-mL-1 in DCM was added dropwise into PEZ solution under vigorous stirring conditions at 25 °C for 24 h to produce PEZ-alt-PEG [28]. The solution was concentrated by rotary evaporation to remove DCM and dialyzed (MWCO 14 kDa) against water. Subsequently, PEZ-alt-PEG copolymers were obtained by lyophilization. Yield: 95%.
Finally, PEZ-alt-PEG was dissolved in TFA, followed by the addition of anisole and methanesulfonic acid, After stirring at room temperature for 1.5 h, the solution was diluted with distilled water and washed three times with diethyl ether. The aqueous layer was collected and then neutralized with sodium bicarbonate. Afterward, the sample was transferred into a dialysis tube (MWCO 7 kDa) and dialyzed for two days against a 5.0 wt% NaHCOg aqueous solution and subsequently five days against distilled water. The dialyzed product was then lyophilized to obtain the final product MPLL-alf-PEG. Yield: 91 %
The nuclear magnetic resonance (NMR) spectra of P EZ and PEZ-alt-PEG were da termined on a Bruker DMX 400 MHz spectrometer (Bruker, Karlsruhe, Germany) using DMSO-M as solvent, while MPLL-alt-PEG was characterized with D2 O as solvent. The molecular weights of various c op polymers were determined by gel permeation chromatography (GPC) systems (Waters, Milford, USA) equipped with a Waters 51 5 HPLC pump and a Waters 2414 refractive index detector. For the characterization of PEZ and PEZ-alt-PEG, samples were analyzed with dimethylformamide (DMF) as the eluent using linear poly (methyl methacrylates) (PMMAs) as standards. For the characterization of MPLL-alt-PEG, the sample was meas and using a 0.50 M H Ac-N an Ac buffer (pH 4.5) as the eluent and linear PEG as st standard.
2.3.Preparation and Characterization drug-Loaded MPLL-alt-PEG Micelles
To evaluate the encapsulation of small anionic molecules by MPLL-alt-PEG, a given amount of MO was added into an MPLL-alt-PEG solution in water (10 mL) and stirred for 5 min. The obtained solution was then dialyzed (MWCO 50 kDa) against water under sink conditions to remove free guest molecules. The solution with an equivalent amount of free MO was selected as the control for dialysis under the same conditions. When the MO solution in the control group was dialyzed to colorless, MO/MPLL-alt-PEG micelles were obtained, and the amounts of MO in the dialysate was determined by UV-vis spectrophotometer. A standard calibration curve was used for quantification of MO with linear correlation coefficient (r2) > 0.99 in the concentration range of 0.1-6 mg-L-1, and the drug loading capacity was calculated as the mass percentage of MO to polymer.
To evaluate the encapsulation of anionic biomacromolecules by MPLL-alt-PEG, IFN was labeled by FITC following the reported method [31]. Subsequently, the MPLL-alt-PEG and FITC-labelled IFN (FITC-IFN) solution were dissolved in a 0.01 M phosphate buffer (pH 7.4) at a concentration of 2 mg-mL-1, respectively. 1 mL of the IFN solution was then added dropwise into 5 mL of polymer solution under sonication for 5 min on a sonicator (KQ-200 KDE, Kunshan Ultrasonic Instrument Co.,
Kunshan, China, 40 kHz, 100 W), and then dialyzed (MWCO 100 kDa) against 0.01 M phosphate buffer (pH 7.4) under sink condition. The dialysis of free FITC-IFN in a 0.01 M phosphate buffer (pH 7.4) was also conducted as a control. When the free IFN in the control experiment had been completely removed, the amount of FITC-IFN in the outer dialysate was measured by a Spectramax Gemini XS microplate fluorometer (Molecular Devices Cooperation, Sunnyvale, CA, USA) using a standard calibration curve of FITC-IFN, and then that value was subtracted from the total amount of added FITC-IFN to calculate the loaded amount of FITC-IFN. The drug loading capacity was calculated as the mass percentage of FITC-IFN to polymer.
The morphologies of formed micelles were visualized with a JEOL JEM-1400 transmission electron microscope at an operating voltage of 100 kV (JEOL, Tokyo, Japan). The particle sizes and zeta potentials of both polymers and micelles were recorded at 25 °C using a Zetasizer Nano ZS90 (Malvern Instruments Ltd., Worcestershire, UK). The solutions were diluted to 1 mg-mL-1 using a 0.01 M phosphate buffer (pH 7.4) before testing.
2.4.pH-Sensitive Release off from Micelle
FITC-IFN-loaded micelle solution (5 mL) was transferred into a dialysis bag (MWCO 100 kDa), and then the bag was placed into 150 mL of a 0.02 M phosphate buffer (pH 7.4), 0.02 M HAc-NaAc buffer (pH 5.5), or 0.02 M HAc-NaAc buffer (pH 4.5). The polymer/FITC-IFN complex solution was prepared as described in Section 2.3. At predetermined time intervals, a given volume of release media was withdrawn and replenished with an equal volume of fresh buffer solution. The amounts of released IFN were determined by the microplate fluorometer, the cumulative amount of released drug was calculated, and the percentages of drug released were plotted against time. All of the experiments were carried out in duplicate.
2.5.Measurement of Minimal Inhibitory Concentration (MIC)
The antimicrobial activities of the polymers were investigated against Gram-positive MRSA and Gram-negative P. aeruginosa using the broth microdilution method. Bacterial cells were incubated in a Mueller Hinton Broth (MHB) medium at 37 °C under constant shaking at 300 rpm to reach the mid-logarithmic growth phase. The microbial suspensions were diluted and adjusted to 2 X 105 colony forming units (CFU)-mL-1. Polymers were subjected to a series of two-fold dilutions using MHB to bacteria concentrations ranging from 60 to 0.2 ^M. A total of 50 of each diluted polymer solution was added to 96-well microplates, followed by the addition of 50 rL of bacterial suspensions, to give a final inoculum of 1 X 105 CFU-mL-1. After being incubated at 37 °C for 18 h, the optical density absorbance at 600 nm was measured with a microplate reader (Synergy HT, BioTek Instruments Inc., Winooski, VT, USA). Bacterial cells without polymer treatment and polymer solutions without bacteria addition were used as positive and negative controls, respectively. The MIC was defined as the lowest concentration that inhibited bacterial growth after 18 h.
2.6.Hemolysis Assay
Freshly mouse blood was treated with K2EDTA, washed with a phosphate buffer solution (PBS) three times and then centrifuged at 1000 g for 10 min to separate erythrocytes. Then, the red blood cells were obtained and diluted to a final concentration of 5% (v/v) with PBS (pH 7.4). 50 rL of polymer solutions at different concentrations were mixed with an equal volume of red blood cell suspensions on a 96-well microplate followed by incubation for 1hat37 °C. After centrifugation at 1000X g for 10 min, aliquots (30 RL) of the supernatant were transferred to another 96-well microplate and diluted with 100 RL of PBS.
The release of hemoglobin from erythrocyte suspensions was monitored by absorbance measurement at 540 nm using a Biotek Synergy TH microplate reader. Absorbance of red blood cells lysed with 2% Triton X-100 was used as the positive control and taken to be 100% hemolytic, while untreated erythrocyte suspension in the PBS solution was used as a negative control. Meanwhile, a control sample of polymer solution was applied to rule out polymer absorption interference (false-positive results), and the existence false-negative results were also checked according to the reported method [32]. Each test was repeated three times. The percentage of hemolysis was calculated according to the following Equation (1):
Hemolysis (%)=[(久 sample — A negative)/0 positive —久 negative)] X 100% (1) where A sample represents the absorbance of the treated sample, ^negative represents the absorbance of the negative control, and Apositive represents the absorbance of the positive control.
All animal experiments in this research were carried out according to the Guidelines for the Care and Use of Laboratory Animals and were approved by the Animal Ethics Committee of Shantou University Medical College (SUMC2016-200, 5 December 2016).
2.7.Imaging Study of microorganisms
Bacteria cells were grown and mixed with polymer at 2X MIC using a protocol similar to the MIC determination. After 90 min incubation at 37 °C, bacteria cells were collected, washed twice with PBS (5000X g, 10 min), and fixed with an equal volume of 2.5% glutaraldehyde solution for 4 h at 4 °C. Bacteria cells were rinsed in a 0.1 M PBS (5000X g, 10 min), and further fixed in 1% osmic acid (4 °C for 2 h). After rinsing in 0.1 M PBS, cells were dehydrated through a graded series of ethanol (30-100%).
For observation by field-emission scanning electron microscopy (FE-SEM), the dehydrated samples were transferred to a 1:1 mixture of ethanol and isoamyl acetate for 30 min, and then to absolute isoamyl acetate for 1 h. The samples were critical point-dried, sputter-coated with a thin layer of gold, and analyzed with FE-SEM (SU8010; Hitachi, Tokyo, Japan).
For observation by transmission electron microscopy (TEM), the dehydrated samples were transferred to acetone for 20 min, to a 1:1 mixture of acetone and Spurr resin for 1 h, and then to a 3:1 mixture of resin and absolute acetone for 3 h. Finally, the samples were transferred to Spurr resin and incubated overnight. Sections having thickness of 70-90 nm were obtained, and post-stained with uranyl acetate and lead citrate for 10 min prior to TEM observations under a TEM setup (H-7650, Hitachi, Tokyo, Japan).
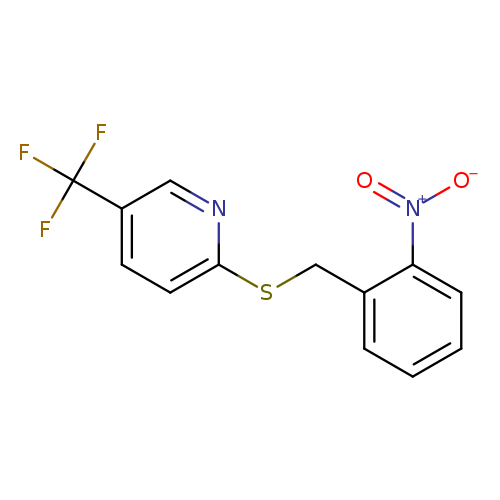
2-{[(2-nitrophenyl)methyl]sulfanyl}-5-(trifluoromethyl)pyridineCatalog No.:AA019UOX CAS No.:1118787-26-0 MDL No.:MFCD11857928 MF:C13H9F3N2O2S MW:314.2830 |
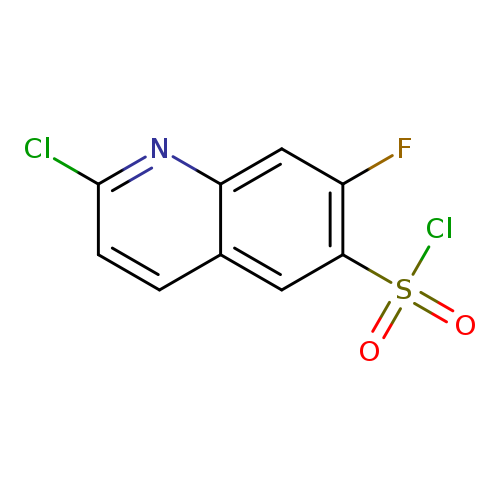
2-Chloro-7-fluoroquinoline-6-sulfonyl chlorideCatalog No.:AA019SDZ CAS No.:1118787-31-7 MDL No.:MFCD11858140 MF:C9H4Cl2FNO2S MW:280.1030 |
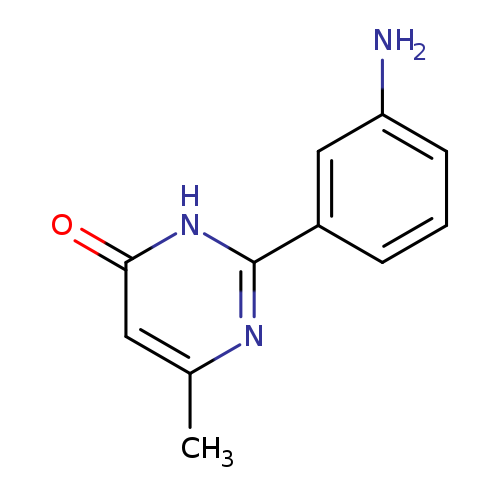
2-(3-aminophenyl)-6-methyl-3,4-dihydropyrimidin-4-oneCatalog No.:AA019UA5 CAS No.:1118787-36-2 MDL No.:MFCD11839789 MF:C11H11N3O MW:201.2245 |
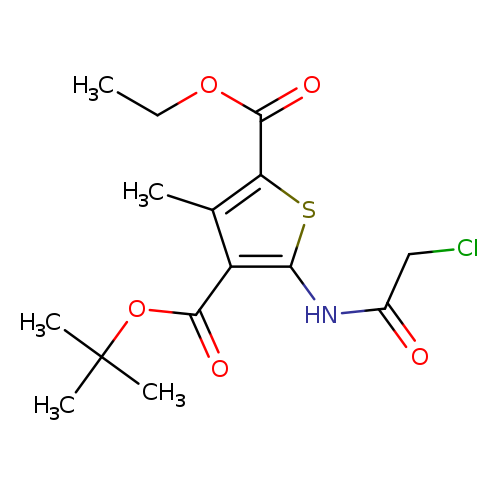
4-tert-butyl 2-ethyl 5-(2-chloroacetamido)-3-methylthiophene-2,4-dicarboxylateCatalog No.:AA019UAG CAS No.:1118787-39-5 MDL No.:MFCD11839804 MF:C15H20ClNO5S MW:361.8410 |
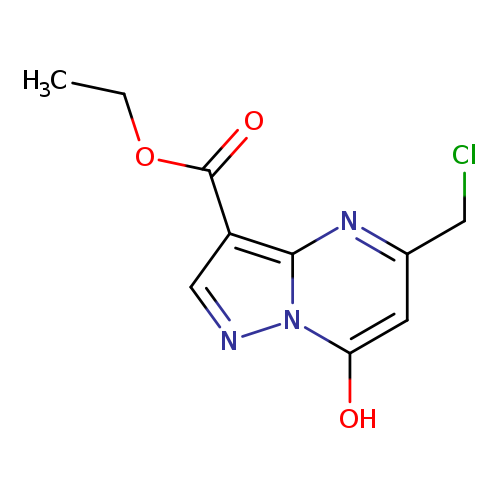
ethyl 5-(chloromethyl)-7-hydroxypyrazolo[1,5-a]pyrimidine-3-carboxylateCatalog No.:AA019UBN CAS No.:1118787-44-2 MDL No.:MFCD11839858 MF:C10H10ClN3O3 MW:255.6577 |
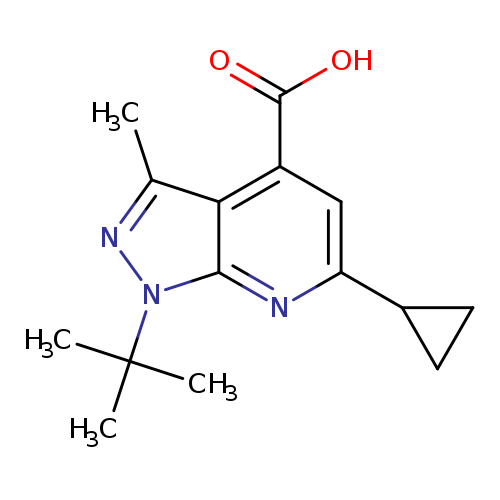
1-tert-butyl-6-cyclopropyl-3-methyl-1H-pyrazolo[3,4-b]pyridine-4-carboxylic acidCatalog No.:AA019UBO CAS No.:1118787-47-5 MDL No.:MFCD11839859 MF:C15H19N3O2 MW:273.3303 |
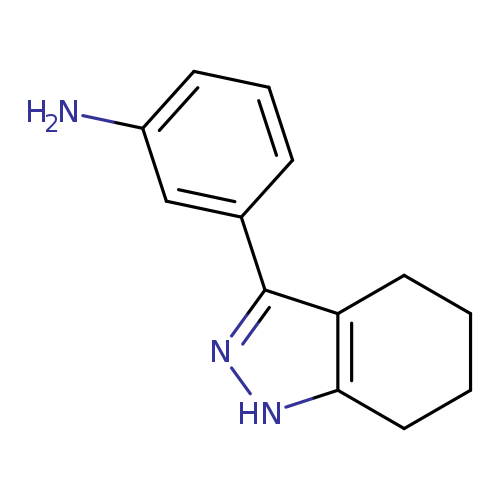
3-(4,5,6,7-tetrahydro-1H-indazol-3-yl)anilineCatalog No.:AA019UC5 CAS No.:1118787-56-6 MDL No.:MFCD11857816 MF:C13H15N3 MW:213.2783 |
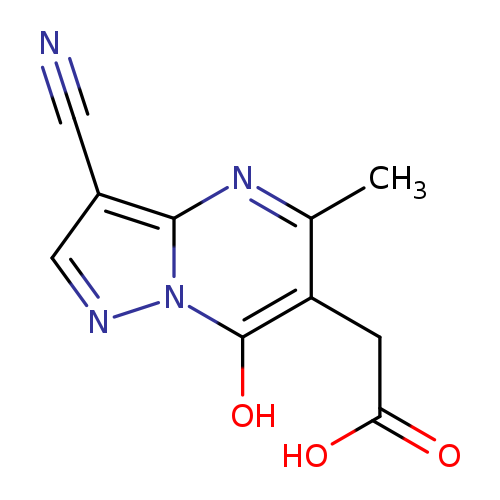
2-{3-cyano-7-hydroxy-5-methylpyrazolo[1,5-a]pyrimidin-6-yl}acetic acidCatalog No.:AA019UD2 CAS No.:1118787-65-7 MDL No.:MFCD11839861 MF:C10H8N4O3 MW:232.1955 |
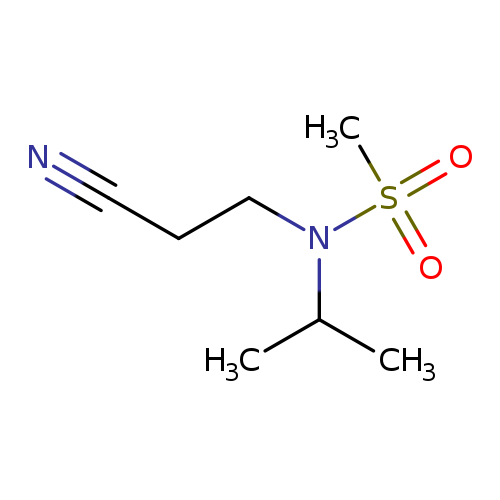
N-(2-cyanoethyl)-N-(propan-2-yl)methanesulfonamideCatalog No.:AA019UD3 CAS No.:1118787-68-0 MDL No.:MFCD11839862 MF:C7H14N2O2S MW:190.2633 |
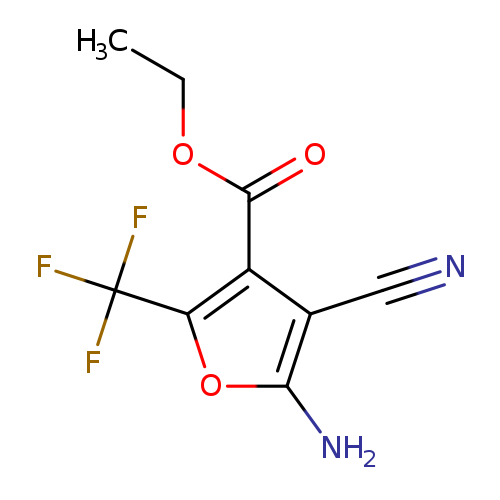
ethyl 5-amino-4-cyano-2-(trifluoromethyl)furan-3-carboxylateCatalog No.:AA019UEZ CAS No.:1118787-87-3 MDL No.:MFCD11857839 MF:C9H7F3N2O3 MW:248.1587 |
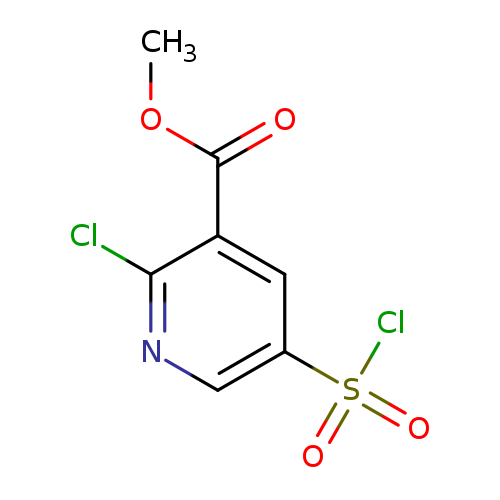
Methyl 2-chloro-5-(chlorosulfonyl)pyridine-3-carboxylateCatalog No.:AA019UF7 CAS No.:1118787-91-9 MDL No.:MFCD11857840 MF:C7H5Cl2NO4S MW:270.0899 |
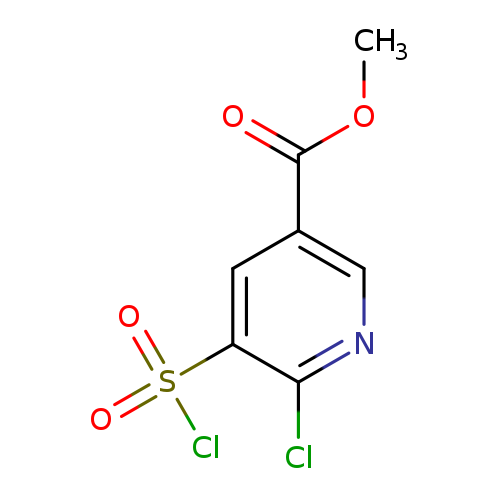
Methyl 6-chloro-5-(chlorosulfonyl)pyridine-3-carboxylateCatalog No.:AA019UF8 CAS No.:1118787-95-3 MDL No.:MFCD11857841 MF:C7H5Cl2NO4S MW:270.0899 |
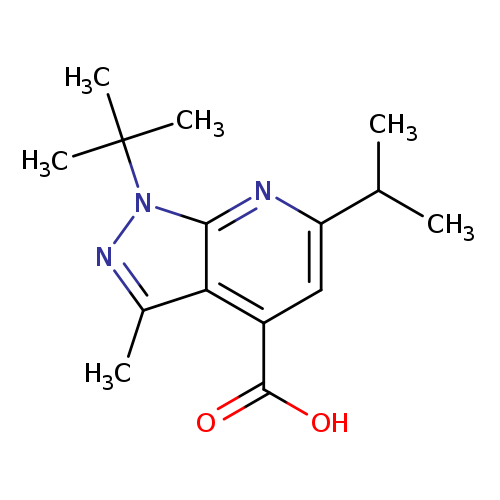
1-tert-butyl-3-methyl-6-(propan-2-yl)-1H-pyrazolo[3,4-b]pyridine-4-carboxylic acidCatalog No.:AA019UFD CAS No.:1118788-02-5 MDL No.:MFCD11840106 MF:C15H21N3O2 MW:275.3461 |
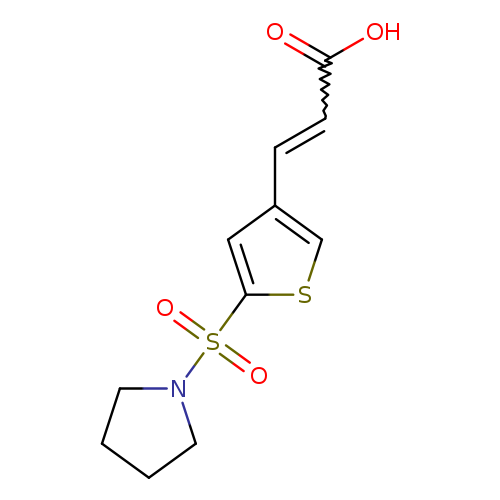
3-[5-(pyrrolidine-1-sulfonyl)thiophen-3-yl]prop-2-enoic acidCatalog No.:AA019UGP CAS No.:1118788-19-4 MDL No.:MFCD11857854 MF:C11H13NO4S2 MW:287.3552 |
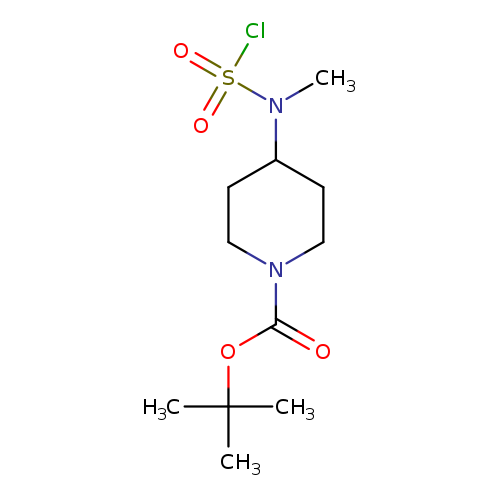
tert-butyl 4-[(chlorosulfonyl)(methyl)amino]piperidine-1-carboxylateCatalog No.:AA019UH8 CAS No.:1118788-30-9 MDL No.:MFCD11857858 MF:C11H21ClN2O4S MW:312.8134 |
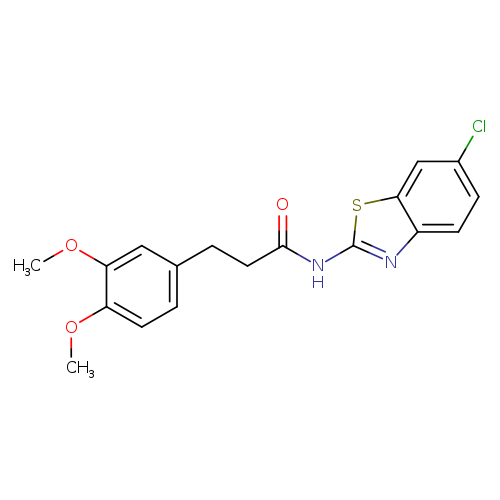
N-(6-Chloro-1,3-benzothiazol-2-yl)-3-(3,4-dimethoxyphenyl)propanamideCatalog No.:AA008TEG CAS No.:1118807-13-8 MDL No.:MFCD08483062 MF:C18H17ClN2O3S MW:376.8572 |
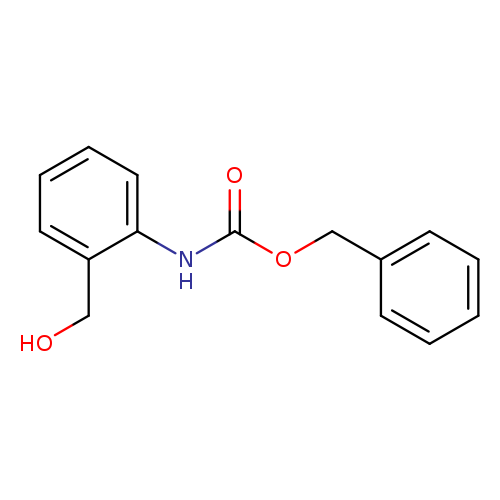
Benzyl n-[2-(hydroxymethyl)phenyl]carbamateCatalog No.:AA009KMX CAS No.:111881-64-2 MDL No.:MFCD11616717 MF:C15H15NO3 MW:257.2845 |
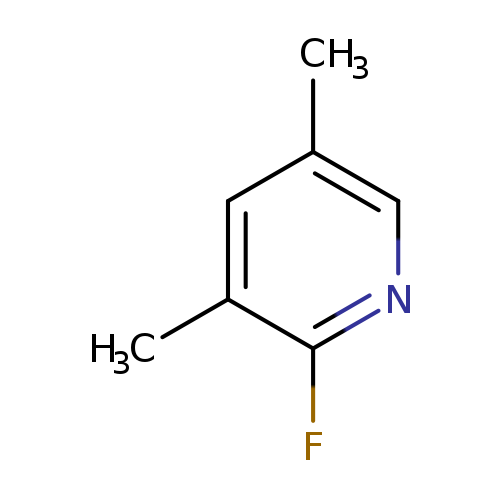
2-Fluoro-3,5-dimethylpyridineCatalog No.:AA008XQP CAS No.:111887-71-9 MDL No.:MFCD11044345 MF:C7H8FN MW:125.1435 |
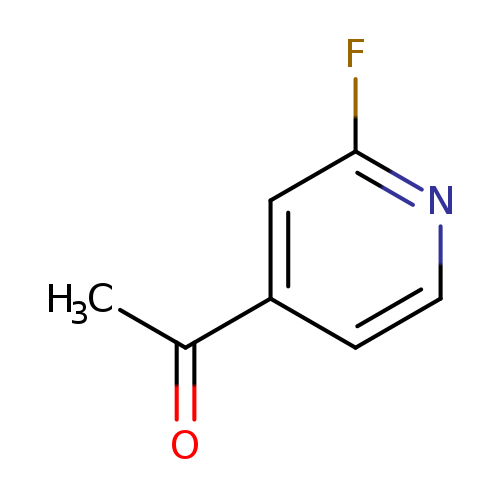
1-(2-Fluoropyridin-4-yl)ethanoneCatalog No.:AA008RBV CAS No.:111887-72-0 MDL No.:MFCD09263924 MF:C7H6FNO MW:139.1270 |

Chloro(1,5-cyclooctadiene)[4,5-dimethyl-1,3-bis(2,4,6-trimethylphenyl)imidazol-2-ylidene] iridium(I)Catalog No.:AA01FR47 CAS No.:1118917-09-1 MDL No.: MF: MW: |
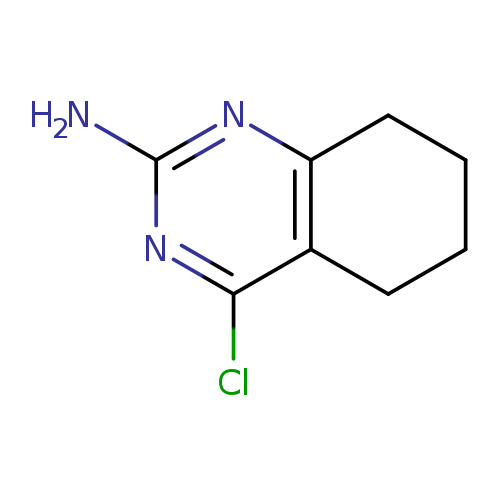
4-Chloro-5,6,7,8-tetrahydroquinazolin-2-amineCatalog No.:AA00HC46 CAS No.:111896-77-6 MDL No.:MFCD13689131 MF:C8H10ClN3 MW:183.6381 |
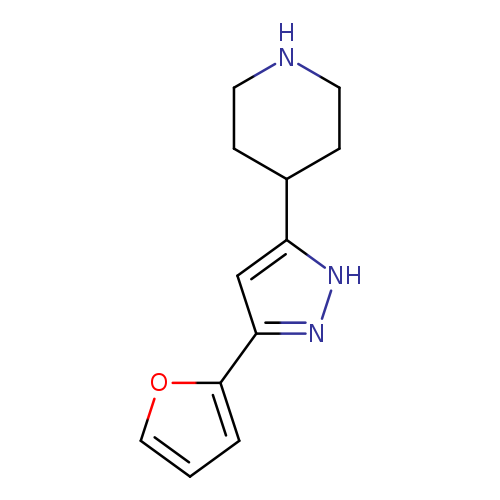
4-(5-(Furan-2-yl)-1H-pyrazol-3-yl)piperidineCatalog No.:AA00IRBQ CAS No.:111897-11-1 MDL No.:MFCD00665128 MF:C12H15N3O MW:217.2670 |
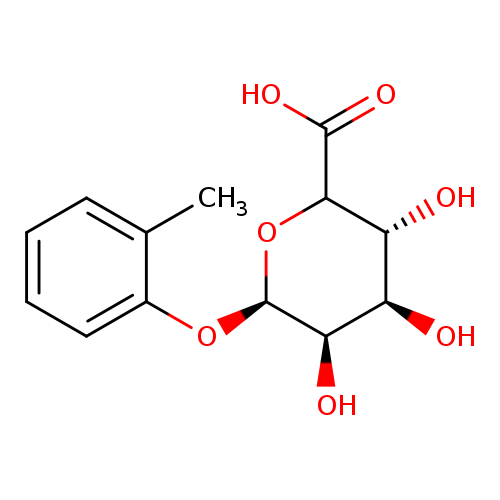
o-Cresol α-D-GlucuronideCatalog No.:AA007VAV CAS No.:111897-99-5 MDL No.:MFCD18252342 MF:C13H16O7 MW:284.2619 |
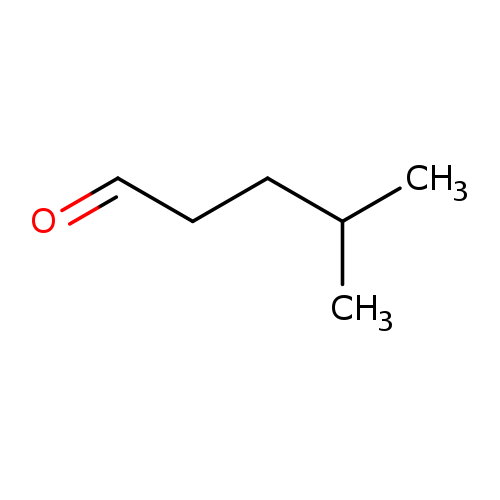
4-MethylvaleraldehydeCatalog No.:AA00385Y CAS No.:1119-16-0 MDL No.:MFCD00871507 MF:C6H12O MW:100.1589 |
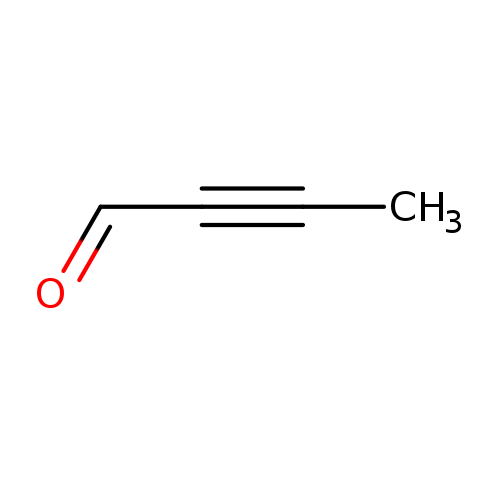
2-Butynal (9CI)Catalog No.:AA0083XS CAS No.:1119-19-3 MDL No.:MFCD12025190 MF:C4H4O MW:68.0740 |
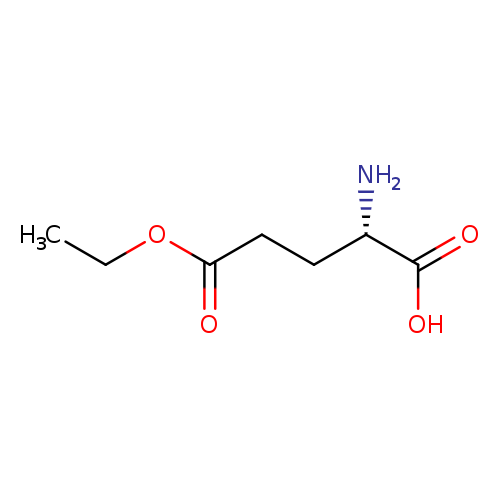
H-Glu(OEt)-OHCatalog No.:AA007DC1 CAS No.:1119-33-1 MDL No.:MFCD00037217 MF:C7H13NO4 MW:175.1824 |
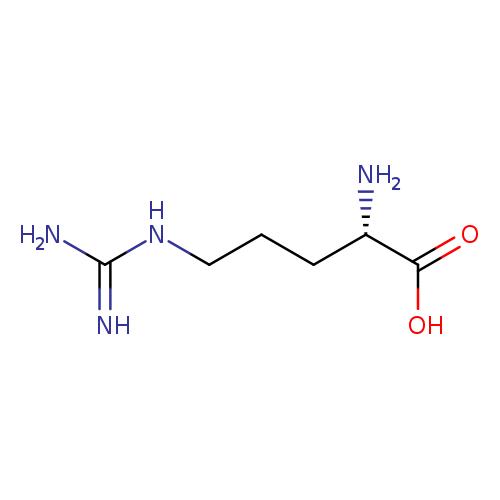
L-Arginine hydrochlorideCatalog No.:AA00HC47 CAS No.:1119-34-2 MDL No.:MFCD00064550 MF:C6H14N4O2 MW:174.2010 |
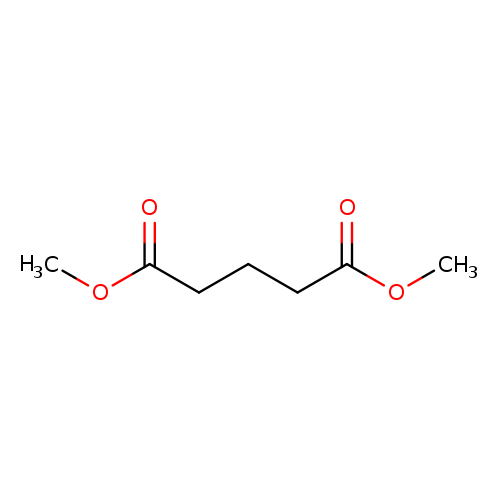
Dimethyl glutarateCatalog No.:AA003QP1 CAS No.:1119-40-0 MDL No.:MFCD00008468 MF:C7H12O4 MW:160.1678 |
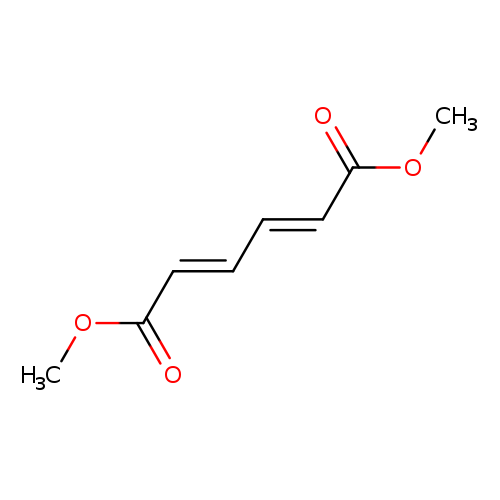
(1E,3E)-1,3-Butadiene-1,4-dicarboxylic acid dimethyl esterCatalog No.:AA008UWY CAS No.:1119-43-3 MDL No.:MFCD00185584 MF:C8H10O4 MW:170.1626 |
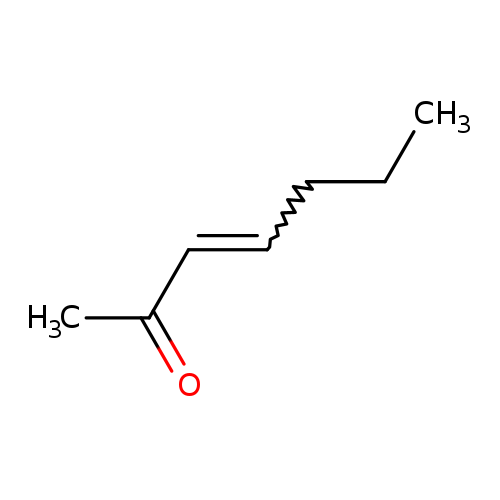
3-Hepten-2-oneCatalog No.:AA003JFO CAS No.:1119-44-4 MDL No.:MFCD00015564 MF:C7H12O MW:112.1696 |
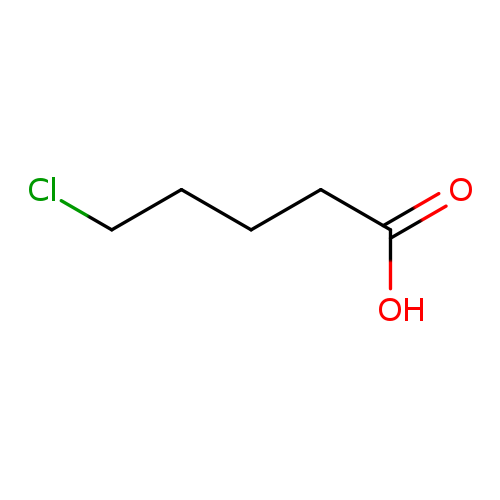
5-Chlorovaleric acidCatalog No.:AA00346M CAS No.:1119-46-6 MDL No.:MFCD00004415 MF:C5H9ClO2 MW:136.5768 |
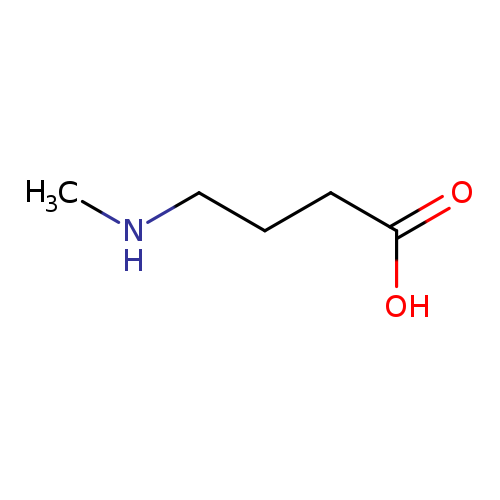
4-(Methylamino)butanoic acidCatalog No.:AA007DB1 CAS No.:1119-48-8 MDL No.:MFCD00044627 MF:C5H11NO2 MW:117.1463 |
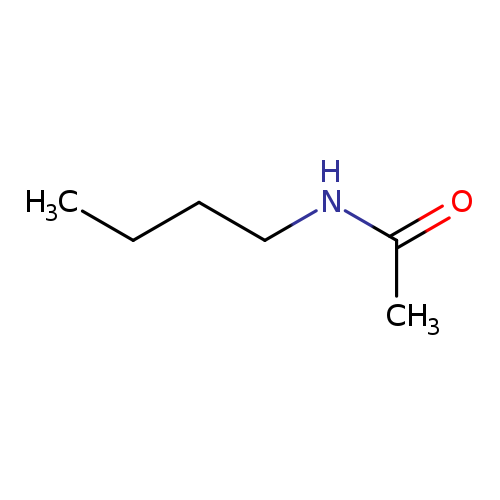
Acetamide, N-butyl-Catalog No.:AA007UZ2 CAS No.:1119-49-9 MDL No.:MFCD00041932 MF:C6H13NO MW:115.1735 |
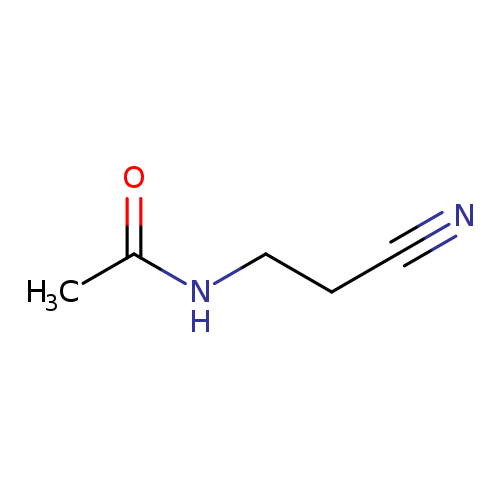
N-(2-Cyanoethyl)acetamideCatalog No.:AA007UZ1 CAS No.:1119-50-2 MDL No.:MFCD00038550 MF:C5H8N2O MW:112.1298 |
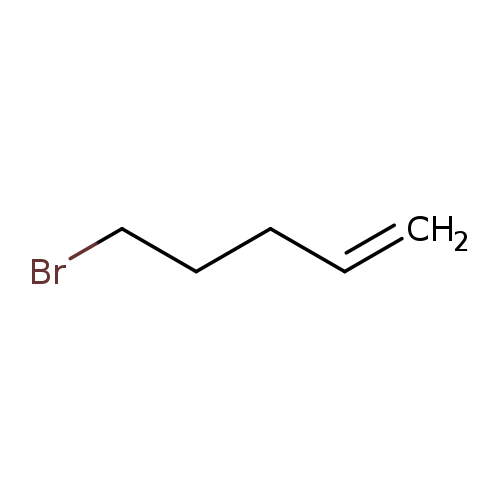
5-Bromo-1-penteneCatalog No.:AA00344F CAS No.:1119-51-3 MDL No.:MFCD00000264 MF:C5H9Br MW:149.0290 |
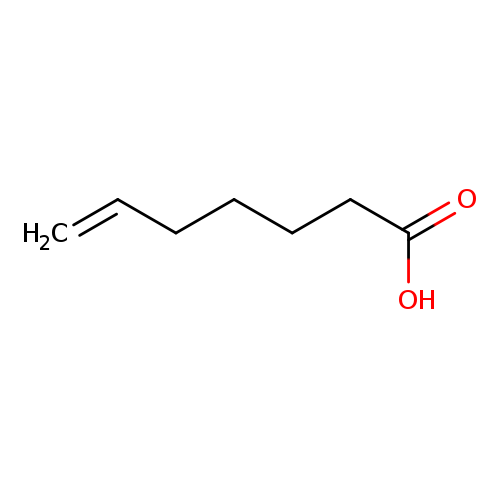
6-Heptenoic acidCatalog No.:AA0037XB CAS No.:1119-60-4 MDL No.:MFCD00014382 MF:C7H12O2 MW:128.1690 |
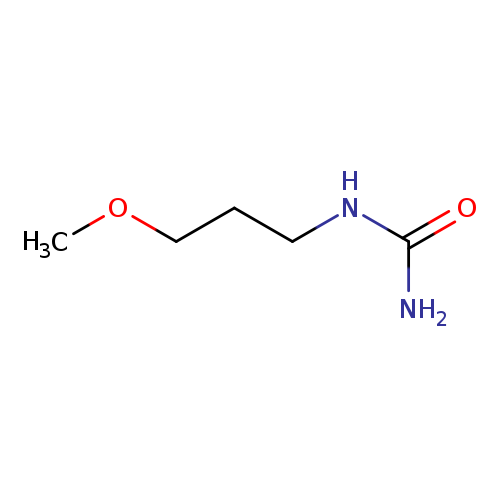
1-(3-Methoxypropyl)ureaCatalog No.:AA007DAX CAS No.:1119-61-5 MDL No.:MFCD03425804 MF:C5H12N2O2 MW:132.1610 |
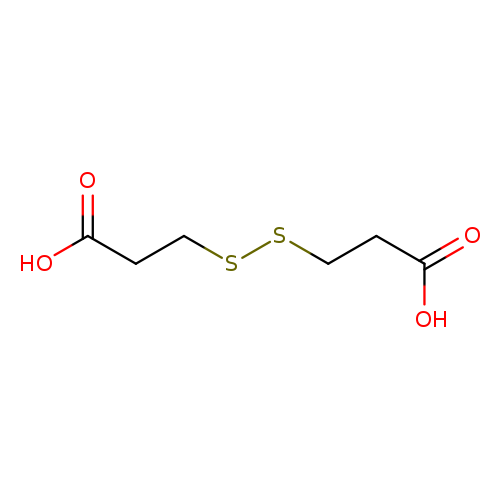
3,3'-Dithiodipropionic acidCatalog No.:AA0033F1 CAS No.:1119-62-6 MDL No.:MFCD00002780 MF:C6H10O4S2 MW:210.2712 |
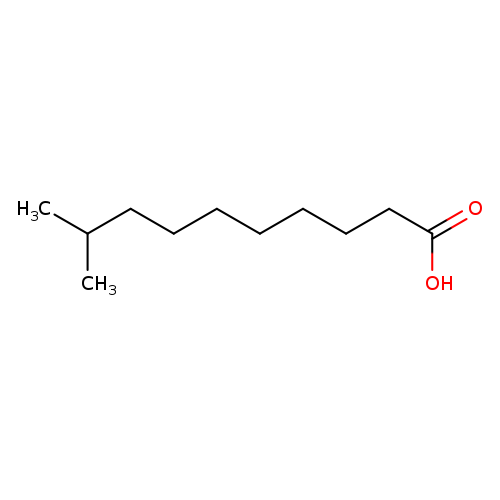
9-Methyldecanoic acidCatalog No.:AA0038WN CAS No.:1119-63-7 MDL No.:MFCD02258677 MF:C11H22O2 MW:186.2912 |
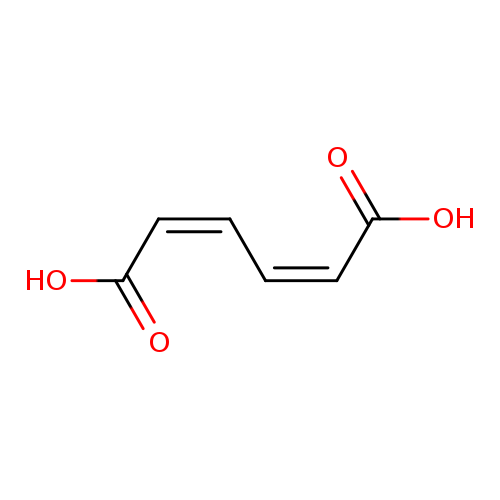
Cis,cis-muconic acidCatalog No.:AA0034JB CAS No.:1119-72-8 MDL No.:MFCD00085266 MF:C6H6O4 MW:142.1094 |
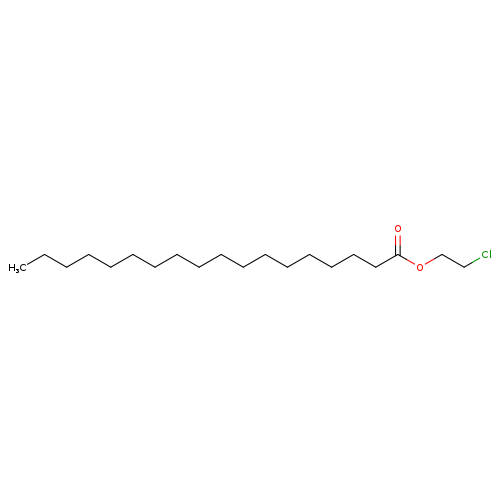
2-Chloroethyl stearateCatalog No.:AA00385U CAS No.:1119-75-1 MDL No.:MFCD06252457 MF:C20H39ClO2 MW:346.9755 |
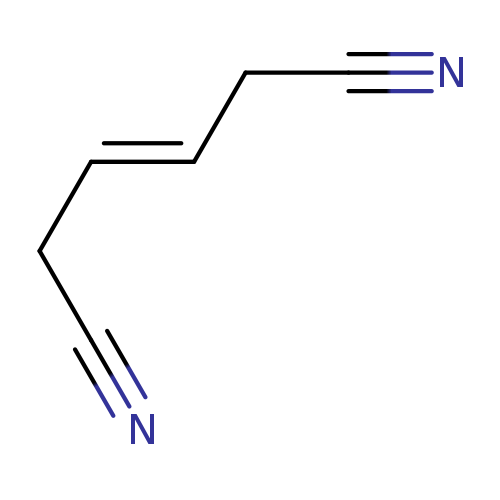
1,4-Dicyano-2-buteneCatalog No.:AA0032K5 CAS No.:1119-85-3 MDL No.:MFCD00001964 MF:C6H6N2 MW:106.1252 |
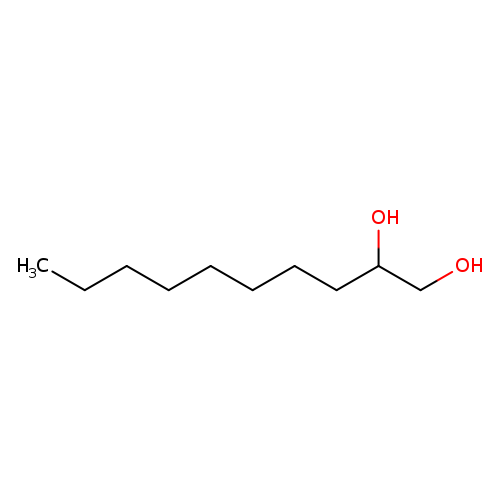
1,2-DecanediolCatalog No.:AA003DAF CAS No.:1119-86-4 MDL No.:MFCD00010739 MF:C10H22O2 MW:174.2805 |
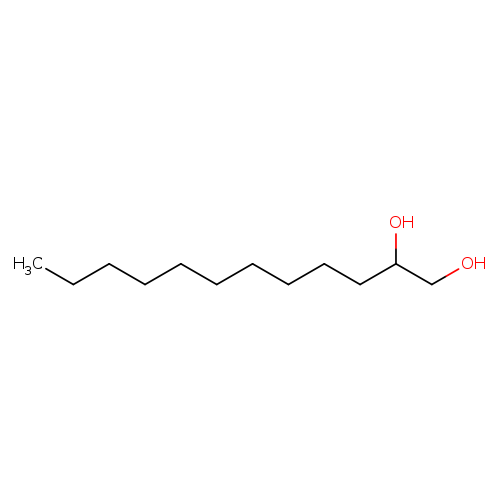
1,2-DodecanediolCatalog No.:AA003PSJ CAS No.:1119-87-5 MDL No.:MFCD00004726 MF:C12H26O2 MW:202.3336 |
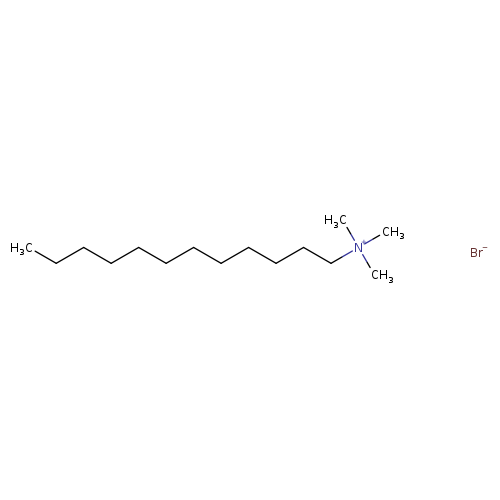
Dodecyltrimethylammonium bromideCatalog No.:AA0083NL CAS No.:1119-94-4 MDL No.:MFCD00011767 MF:C15H34BrN MW:308.3412 |
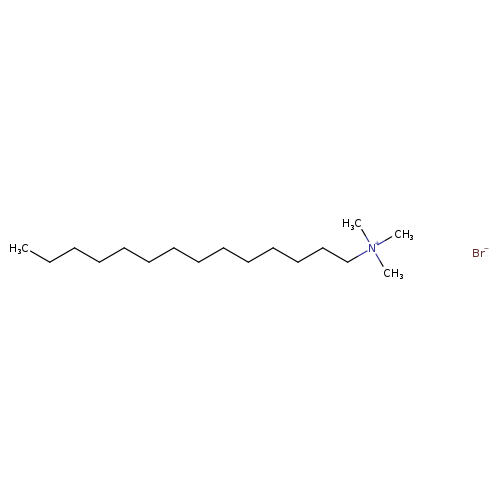
Myristyltrimethylammonium bromideCatalog No.:AA007UV1 CAS No.:1119-97-7 MDL No.:MFCD00011770 MF:C17H38BrN MW:336.3943 |
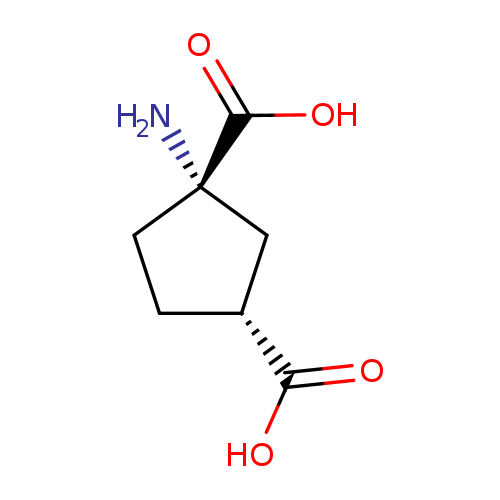
trans-(1S,3R)-ACPDCatalog No.:AA01E60G CAS No.:111900-32-4 MDL No.:MFCD22370015 MF:C7H11NO4 MW:173.1665 |
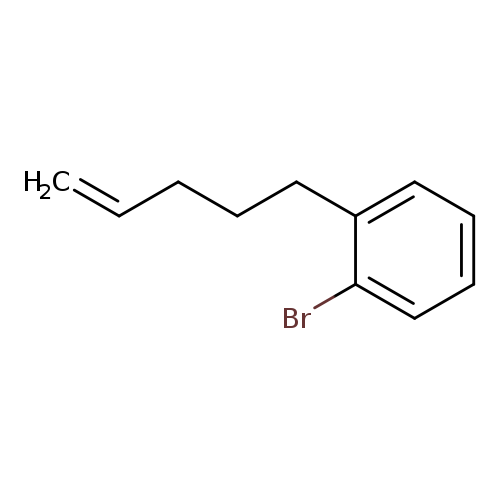
1-bromo-2-(pent-4-en-1-yl)benzeneCatalog No.:AA01BYJW CAS No.:1119036-64-4 MDL No.:MFCD30497736 MF:C11H13Br MW:225.1249 |
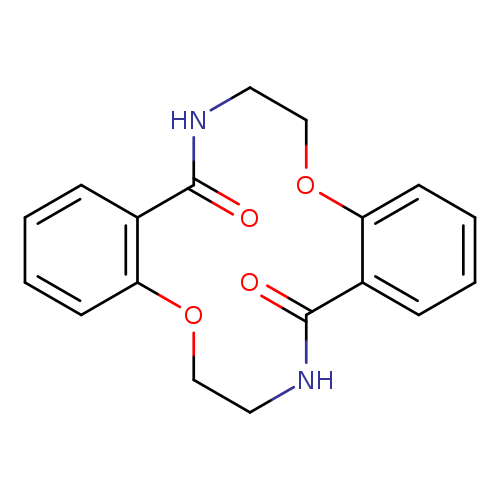
7,8,16,17-Tetrahydro-dibenzo[f,m][1,8,4,11]dioxadiazacyclotetradecine-9,18(6H,15H)-dioneCatalog No.:AA00HC48 CAS No.:111904-37-1 MDL No.:MFCD30834412 MF:C18H18N2O4 MW:326.3465 |
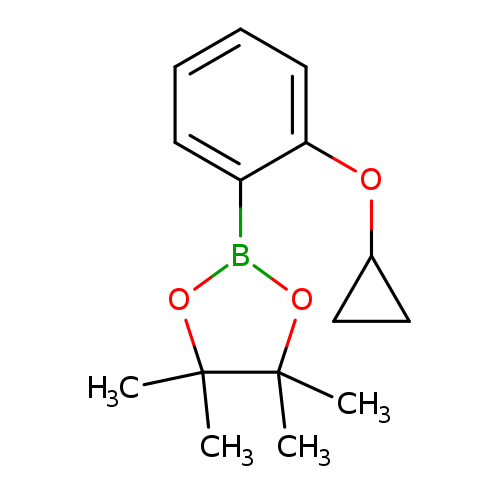
2-(2-Cyclopropoxyphenyl)-4,4,5,5-tetramethyl-1,3,2-dioxaborolaneCatalog No.:AA008U3M CAS No.:1119090-11-7 MDL No.:MFCD16994462 MF:C15H21BO3 MW:260.1364 |
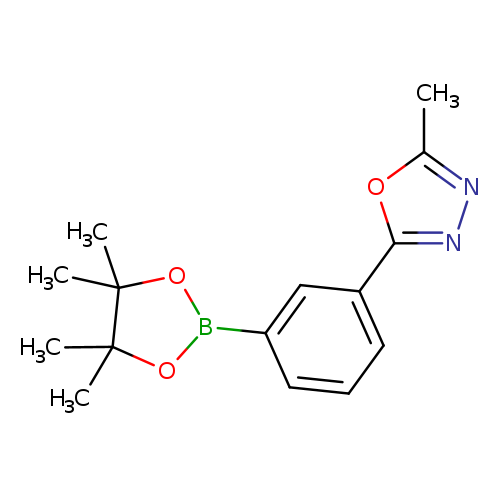
2-Methyl-5-[3-(4,4,5,5-tetramethyl-1,3,2-dioxaborolan-2-yl)phenyl]-1,3,4-oxadiazoleCatalog No.:AA00HC49 CAS No.:1119090-20-8 MDL No.:MFCD18384930 MF:C15H19BN2O3 MW:286.1340 |
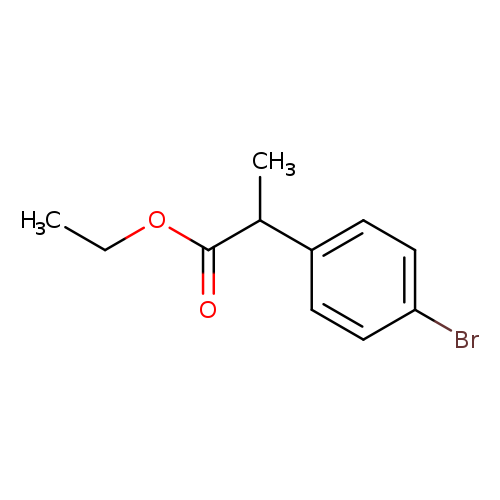
Ethyl 2-(4-bromophenyl)propanoateCatalog No.:AA00HC4B CAS No.:111914-79-5 MDL No.:MFCD18908972 MF:C11H13BrO2 MW:257.1237 |
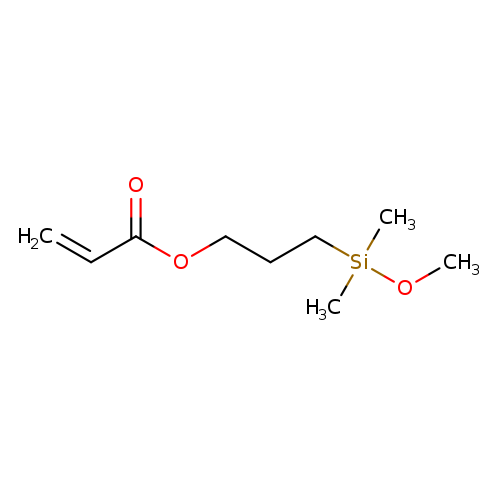
3-(Methoxydimethylsilyl)propyl acrylateCatalog No.:AA007DCL CAS No.:111918-90-2 MDL No.:MFCD00054801 MF:C9H18O3Si MW:202.3229 |
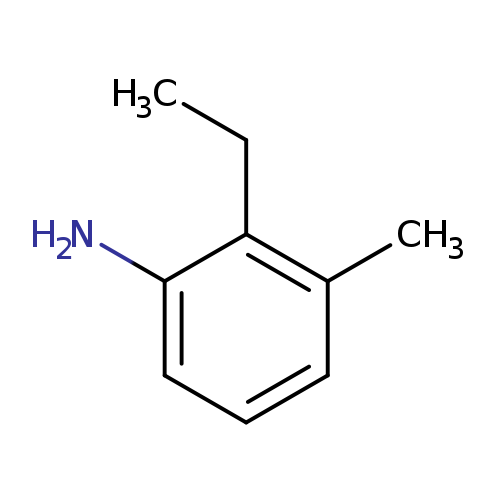
2-ethyl-3-methylanilineCatalog No.:AA01BIL9 CAS No.:111923-32-1 MDL No.:MFCD19205427 MF:C9H13N MW:135.2062 |
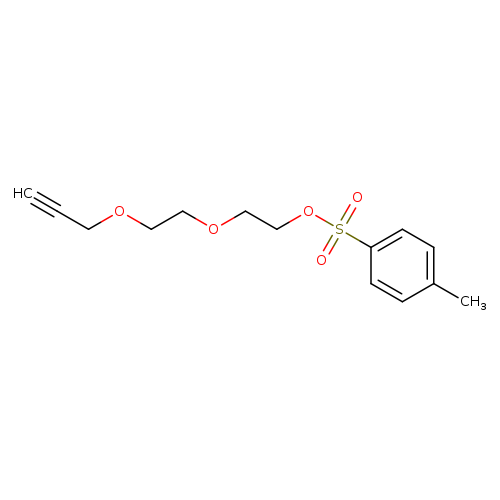
Propargyl-PEG4-TosCatalog No.:AA019DYV CAS No.:1119249-30-7 MDL No.:MFCD20134051 MF:C14H18O5S MW:298.3547 |
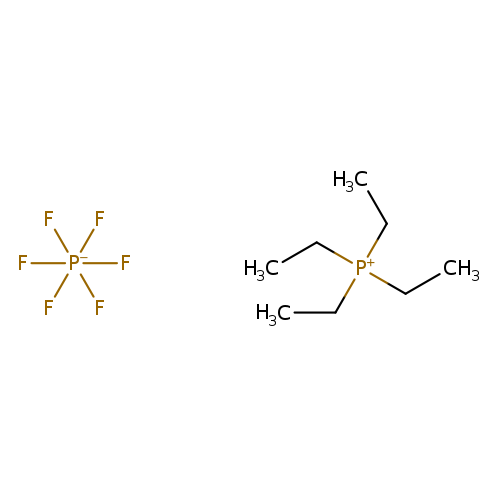
Tetraethylphosphonium hexafluorophosphateCatalog No.:AA003ULL CAS No.:111928-07-5 MDL No.:MFCD01631313 MF:C8H20F6P2 MW:292.1823 |
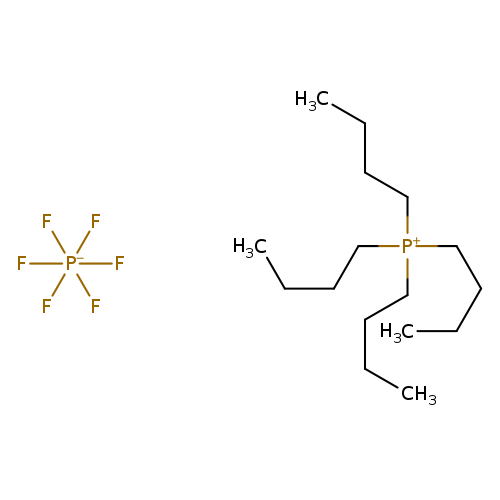
Tetrabutylphosphonium hexafluorophosphateCatalog No.:AA003UKA CAS No.:111928-21-3 MDL No.:MFCD01863114 MF:C16H36F6P2 MW:404.3950 |
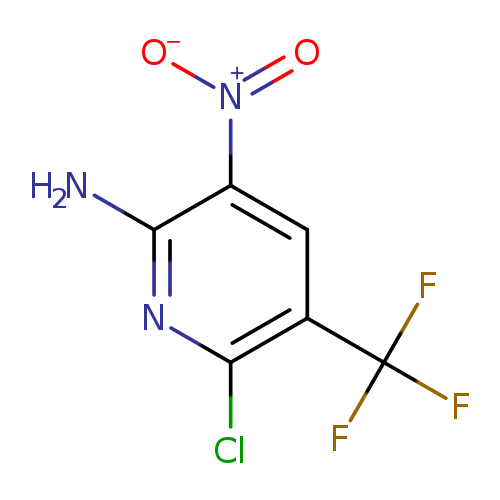
6-Chloro-3-nitro-5-(trifluoromethyl)pyridin-2-amineCatalog No.:AA0095DB CAS No.:111928-64-4 MDL No.:MFCD27987148 MF:C6H3ClF3N3O2 MW:241.5551 |
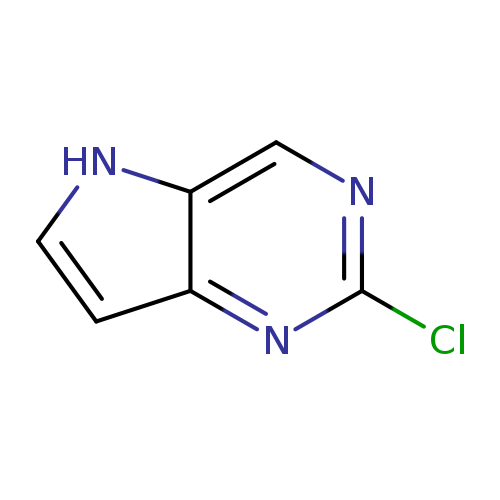
2-Chloro-5H-pyrrolo[3,2-d]pyrimidineCatalog No.:AA00385X CAS No.:1119280-66-8 MDL No.:MFCD11869949 MF:C6H4ClN3 MW:153.5691 |
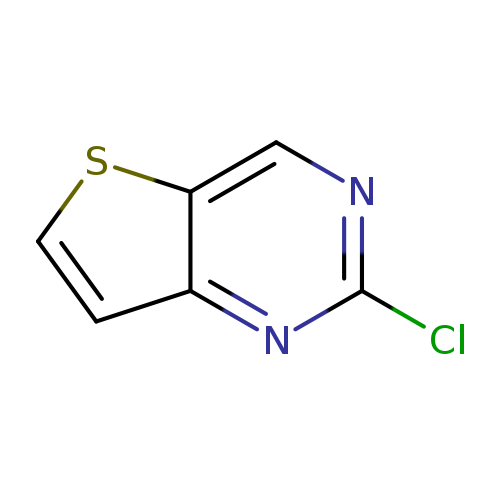
2-Chlorothieno[3,2-d]pyrimidineCatalog No.:AA008S0M CAS No.:1119280-68-0 MDL No.:MFCD12406127 MF:C6H3ClN2S MW:170.6194 |
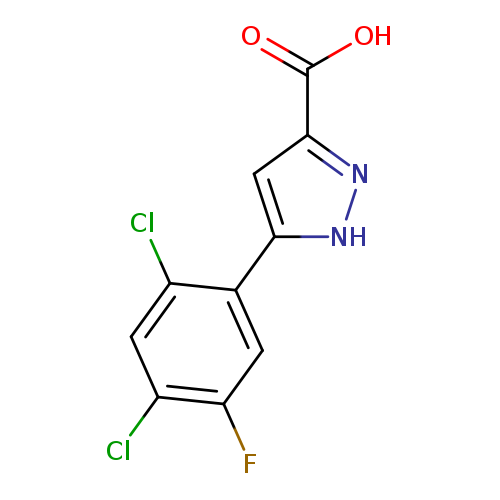
5-(2,4-Dichloro-5-fluorophenyl)-1h-pyrazole-3-carboxylic acidCatalog No.:AA008UQE CAS No.:1119299-75-0 MDL No.:MFCD05864802 MF:C10H5Cl2FN2O2 MW:275.0633 |
Ethyl 2,2-DifluorooctanoateCatalog No.:AA01FF9Y CAS No.:111934-95-3 MDL No.:MFCD15530337 MF:C10H18F2O2 MW:208.2455 |